The distance matrix is selected to best reflect the nature of the data and thus acts as a sort ofįilter or conduit through which data are converted into a form suitable for analysis.Ī consequence of basing analyses on distance matrices is that the resulting axes scores (coordinates) or patternsĪre not independent of one another.
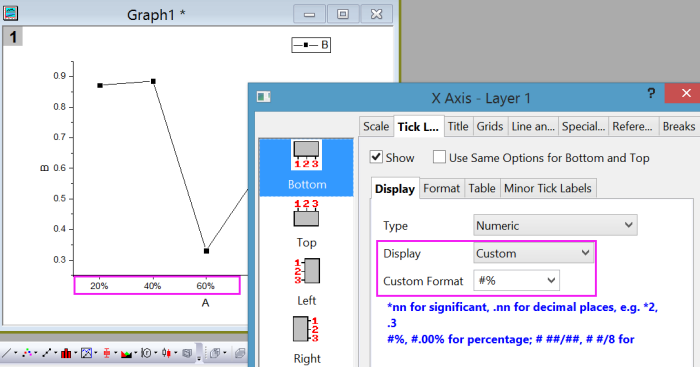
This unrestricted choice of starting distance matrix provides enormous flexibility to be able to handle a wide range ofĭata and issues. (CA/CCA), Q-mode analyses (such as MDS) provide euclidean or arbitrary (where only the order is preserved) ordination distances from any distance or similarity matrix. Matrix algebra is used to arrange objects (sites) so as to preserve either their euclidean distances (PCA/RDA) or $\chi^2$ distances Whereas the starting point of R-mode analyses is typically either a covariance or correlation matrix and In contrast to R-mode analyses (such as principle components analysis), Q-mode analysesĮxplore the patterns between objects (sites) on the basis of pairwise object likenesses (c.f. > set.seed ( 1 ) > x n sp1 sp2 sp3 sp4 sp5 sp6 sp7 sp8 sp9 sp10 X #X colnames (X) rownames (X) X data <- ame ( Sites = factor ( rownames (X), levels = rownames (X)), X) Sites
That extends in a northerly direction over a mountain range. This data set comprises the abundances of 10 species within 10 sites located along a transect To assist with demonstrating Multidimensional Scaling (MDS), we will return to theįabricated species abundance data introduced

